The Clinical Proteomic Tumor Analysis Consortium (CPTAC) project
Overview
The Clinical Proteomic Tumor Analysis Consortium (CPTAC) is a comprehensive and coordinated effort to accelerate understanding of the molecular basis of cancer through the application of robust, quantitative, proteomic technologies and workflows.
The CPTAC analyzes cancer biospecimens from genomics initiatives such as The Cancer Genome Atlas (TCGA) by mass spectrometry to characterize and quantify their constituent proteins or “proteome”. These mass spectrometry data are present in four different file formats: raw, mzML, psm, and mzid. Raw files contain raw mass spectrometry spectra in vendor-specific file formats corresponding to the mass spectrometers used to acquire the spectra. The mzML files are generated by converting these raw files to a HUPO Proteome Standards Initiative (PSI)-compliant format. The psm files report the peptide spectrum match (PSM) data obtained by processing the mzML files. The mzID files were generated by converting the psm files to the HUPO PSI-compliant mzldentML format.
Learn more about the CPTAC data data available on the CGC via this public project.
Access the CPTAC public project
To access the CPTAC public project:
- Click on Public projects from the top navigation bar.
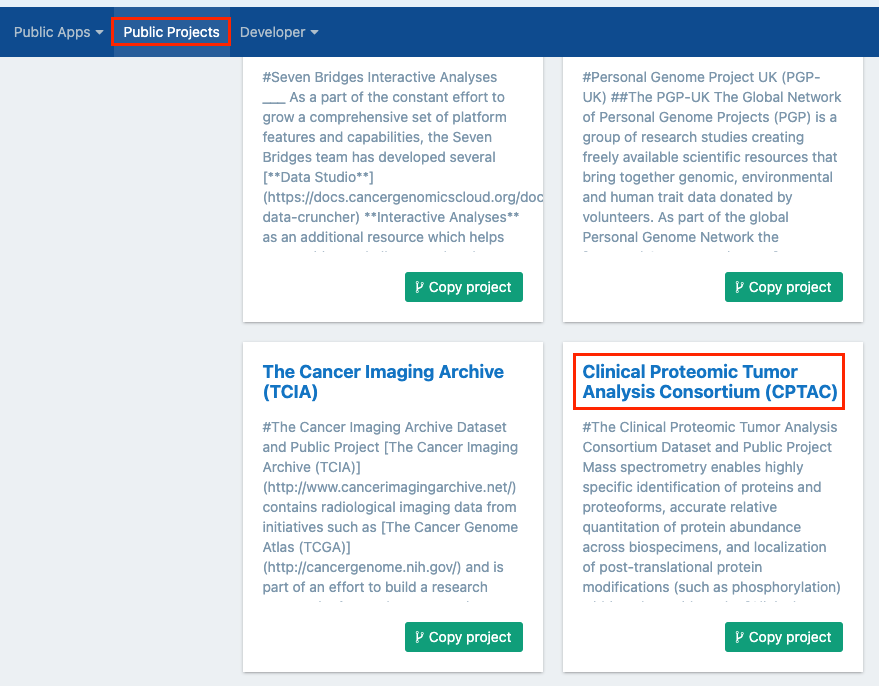
- Select "Clinical Proteomic Tumor Analysis Consortium (CPTAC))" by clicking on its title in the public projects gallery. You'll be taken to the main dashboard of the CPTAC public project.
Use the CPTAC public project
All CGC users automatically have copy permissions for this project. This means that while you cannot upload data or tools to the public project, you can copy the available data to your own projects on the CGC to execute analyses.
You have the options to:
- Copy the entire project - Start from the copied project and add apps to execute analyses on CPTAC data.
- Select and copy a subset of the data to your own project - Use the selected data within your own analyses in your project.
Copy the entire project
- Click Public projects in the top navigation bar.
- Locate the project and click Copy project in the lower right corner.
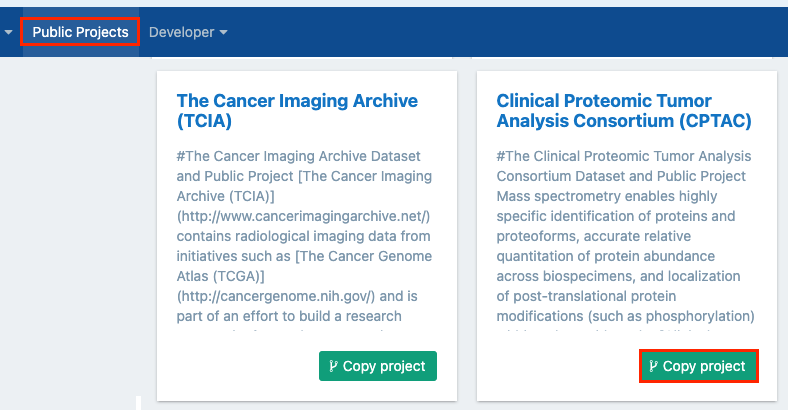
- In the pop-up window, you can name your copy of the project, select a billing group and decide whether the project will contain controlled data..
- Once you've customized the details, click Copy to copy the entire project.
You'll be redirected to the dashboard of your cloned project. Add apps to conduct analyses on the data in your project.
Use a subset of the data
Instead of cloning the entire project, you can choose to select and copy a subset of the data.
- Access the CPTAC public project (see above).
- Click the Files tab. This will take you to the Files page for the CPTAC public project.
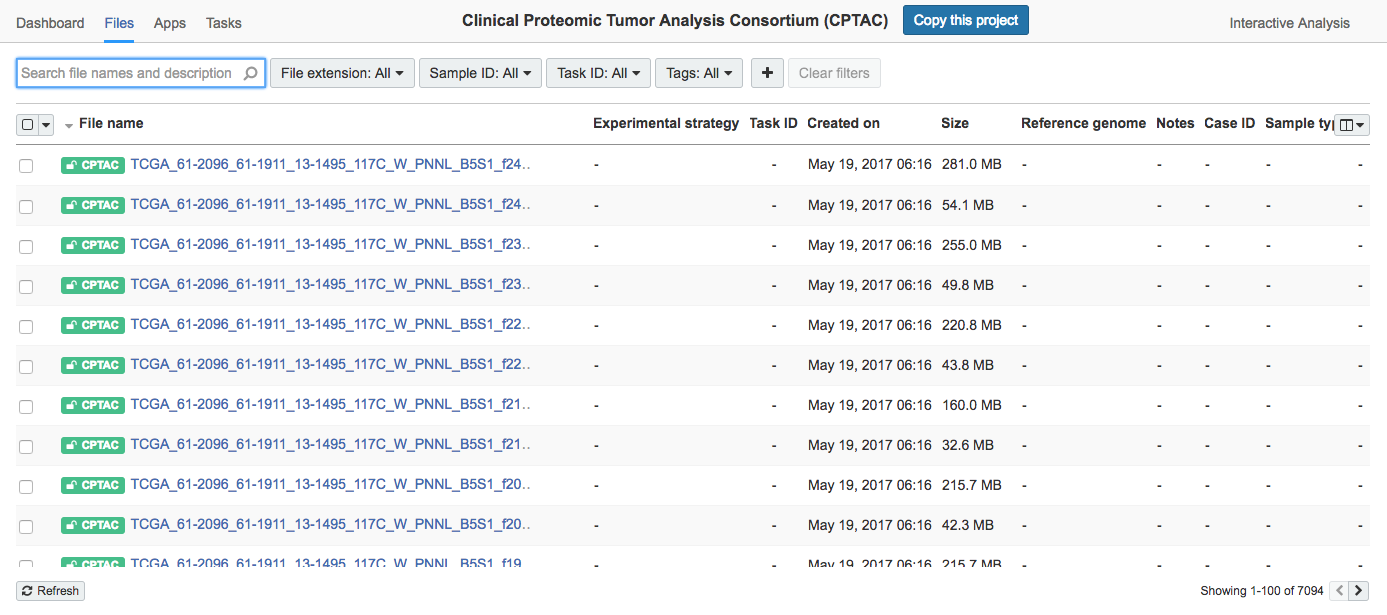
- Filter or search for the desired files.
- Choose specific files by checking the corresponding box by each file name.
- Select as many files as you desire and click Copy to.
- Select your target project from the drop-down menu.
Now, you can start using the CPTAC files you've added to your personal project in your own analysis.
Updated about 3 years ago
