↳ Data Browser query: multiple dataset query
QUERY DATASETS >About the Data Browser > Data Browser query: multiple dataset query
Overview
The Data Browser allows you to query multiple datasets. Once you select the datasets, the entities in the search page will be prioritized based on the number of datasets they appear in. Once you start the search, the Data Browser will check in the selected datasets and give you the results that are found in most of the datasets, while those that no longer apply will be denoted as inactive in the query.
Furthermore, once you select the result and move on to the canvas, as you continue selecting entities, properties, and values, the Data Browser will apply the filters to all datasets that are still relevant to the query and inform you of the datasets that are no longer used (under Active Datasets section).
Finally, when available (the toggle will be enabled), you can apply the intersection for all datasets to narrow down the results to ID properties that are present in all selected datasets. For example, you can apply the intersection to a Case entity and search through information that is related to the same patients across multiple datasets. The only current limitation is that you cannot select TCGA GRCh38 and TCGA Legacy as those are different versions of the same dataset.
Procedure
To start querying multiple datasets:
- Choose Data Browser from the Data menu.
- Select datasets TCGA GRCh38, TARGET GRCh38, and CPTAC.
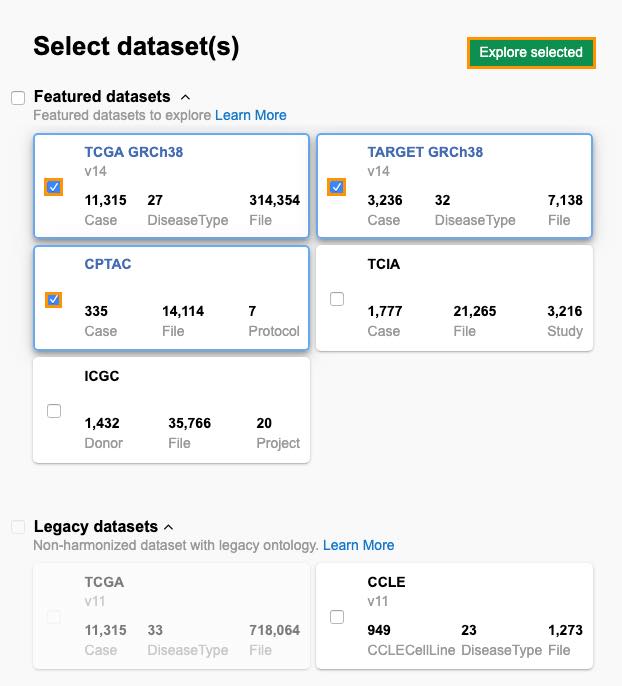
- Click Explore selected.
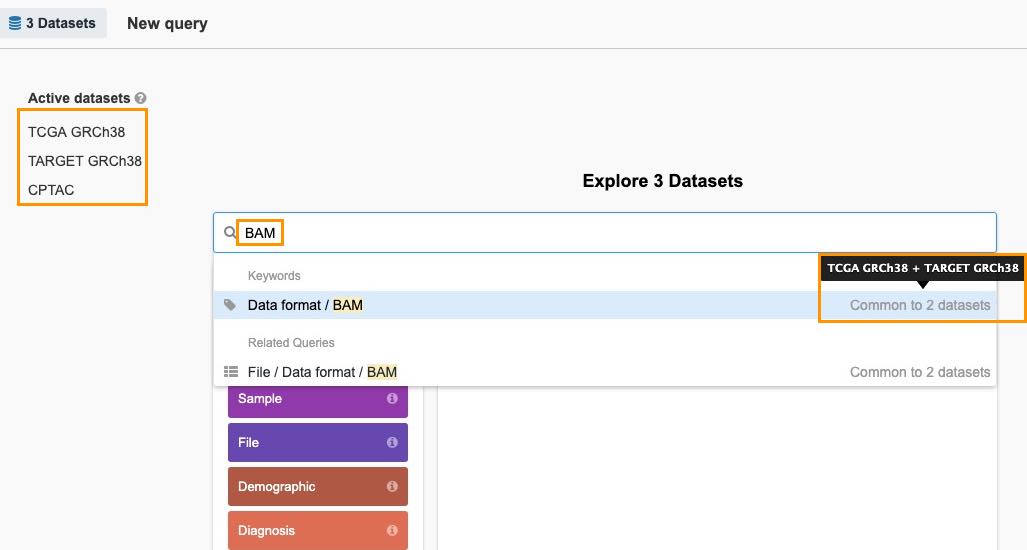
- Enter BAM as a search term and you will see the available results below. The information on how many datasets contain the term is displayed on the right. Hover it to see the dataset names. The list of active datasets on the left will be refreshed as you enter terms to indicate the datasets that are still used in the query i.e. which still contain relevant information based on the entered search terms.
- Click on the result to select it.
- Now enter FEMALE as a term and select the result.
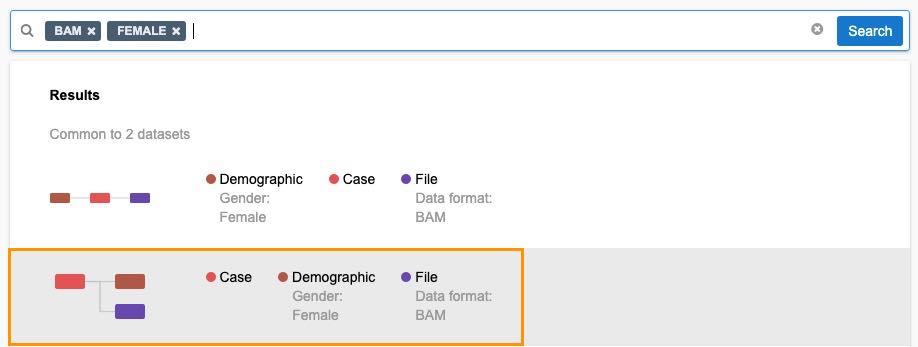
- Click Search.
- Click on the result in the list to continue working with the Data Browser canvas.
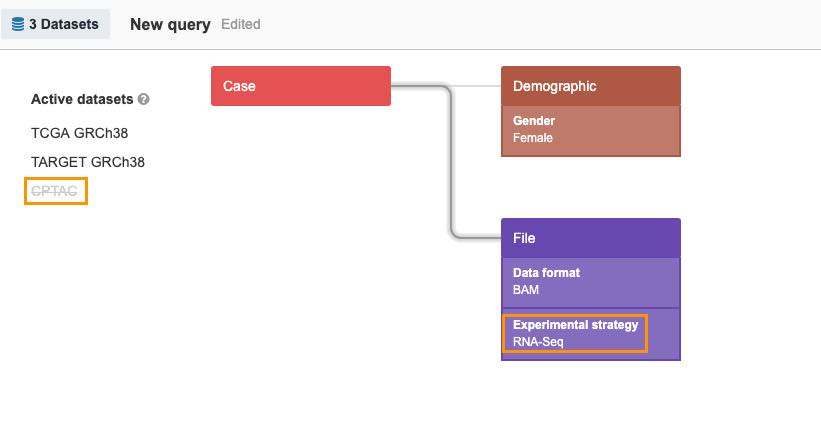
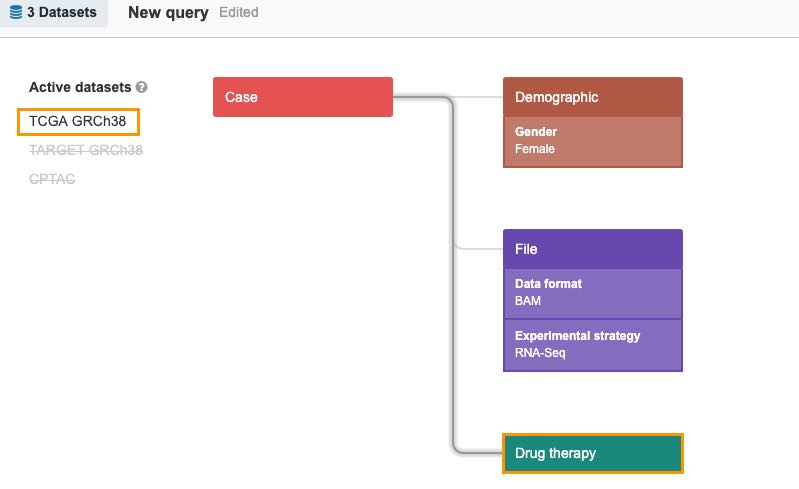
- Add RNA-seq as experimental strategy. As seen in the upper left corner, TCGA GRCh38 and TARGET GRCh38 still contain relevant information and are denoted as active in the query, while CPTAC is still excluded.
- Now, if you add Drug therapy, you will see that the only remaining active dataset in the query is TCGA GRCh38.
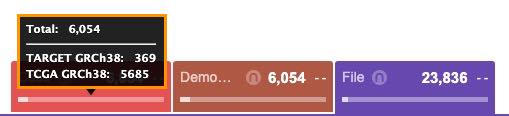
- Hover the bar in the count card to see the distribution of entities across datasets.
Querying datasets with shared data
The Data Browser allows you to query multiple datasets with shared data. The example below will demonstrate the intersection feature which allows selecting files from multiple datasets that apply to the same group of patients (cases).
- Choose Data Browser from the Data menu.
- Select datasets TCGA GRCh38, CPTAC, and TCIA. These datasets share entity instances, i.e. there is a certain number of patients that participated in all of the studies.
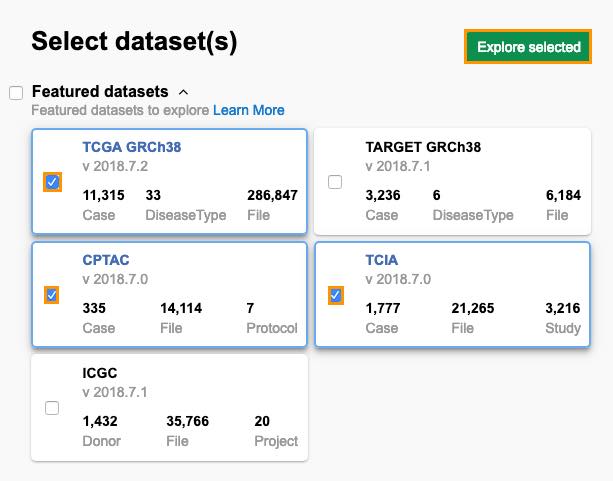
- Click Explore selected.
- Click the Case entity and you will be taken to the Data Browser canvas.
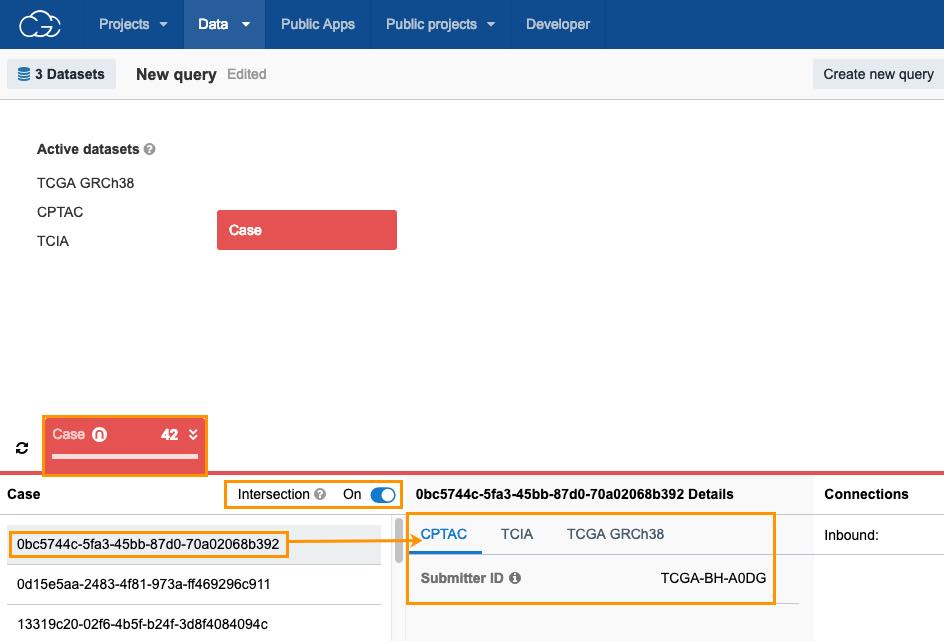
- Click the Intersection toggle to enable the intersection for the selected datasets. The number of cases will be updated (e.g. 42 instead of 11318). To see detailed dataset information for each of the instances, click on an instance under the Case column.
- You can now add files (instances) which belong to multiple datasets, e.g. choose files with the following properties:
a. data format: BAM (from TCGA GRCh38).
b. file type: proteome (from CPTAC).
c. data format: DCM (from TCIA).
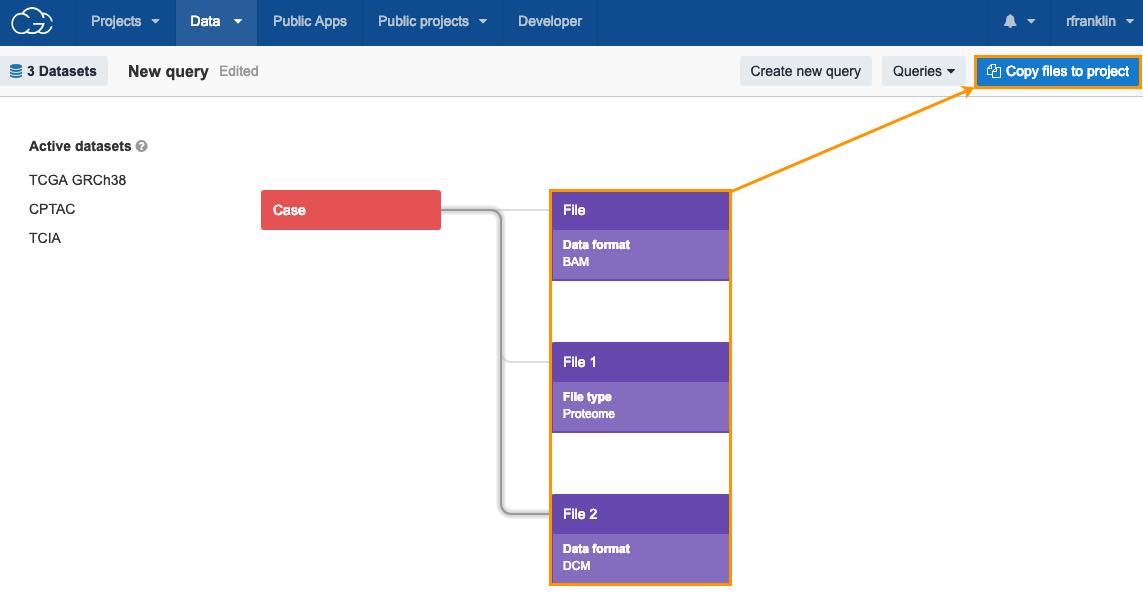
- After adding files with these properties, click Copy files to project to continue with your analysis.
Please bear in mind that multiple datasets can have shared data for more than one entity. In that case the intersection can be applied for one entity at a time.
Updated less than a minute ago
