Personal Genome Project UK (PGP-UK) pilot dataset
The Personal Genome Project UK (PGP-UK) public project on the CGC is made possible through a collaboration between University College London and Seven Bridges. It contains Open Access files from 13 individuals who participated in the PGP-UK pilot study that has been published as a preprint on biorxiv. Through this public project, you will be able to filter the available files according to file type and descriptive metadata and to copy the files of interest into your own project for analysis using your own tools or tools available in the CGC Public Apps gallery.
You don’t need special access or authorization status to use the data available in this project. In fact, any data you copy from this public project into your own projects will not count towards your storage.
These data are made available under the CC-0 license.
WHAT’S CONTAINED IN THE PROJECT?
The Personal Genome Project UK (PGP-UK) pilot data contains 201 files from 13 individuals that were obtained through multi-omics profiling using whole genome sequencing, whole genome bisulphite sequencing, deep and shallow RNA sequencing, and DNA methylation array profiling from saliva or blood.
The PGP-UK public project contains the following distribution of samples and files by experimental strategy.
| Experimental strategy | Number of samples (sample type) | Number of files (file types) |
|---|---|---|
| WGS | 11 (blood) | 55 (fastq, bam, bam.bai, vcf) |
| WXS | 2 (blood) | 8 (fastq, bam, bam.bai, vcf) |
| RNA-Seq (targeted/whole) | 10 (blood) | 50 (fastq, bam, bam.bai) |
| WGBS | 10 (blood) | 40 (fastq.gz, bam, bam.bai) |
| Methylation Array | 11 (blood) + 3 (saliva) | 48 (idat) |
Further information on the samples included in the PGP-UK pilot study can be found in Table 1 of the preprint.
ACCESS THE PGP-UK PUBLIC PROJECT
- Click on Public projects from the top navigation bar.
- Select "Personal Genome Project UK (PGP-UK)" by clicking on its title in the public projects gallery. You'll be taken to the main dashboard of the public project.
USE THE PGP-UK PUBLIC PROJECT
All CGC users automatically have copy permissions for this project. This means that while you cannot upload data or tools to the project, you can copy the available data to your own projects on the CGC and execute analyses there.
You have the options to:
- Copy the entire project - Start from the copied project and add apps to execute analyses on the data.
- Select and use a subset of the data in your own project - Copy the selected data to use within your own analyses.
COPY THE ENTIRE PROJECT
While you cannot directly execute workflows in the PGP-UK public project, you can make a copy of the entire project to perform further analyses.
To copy the entire project:
- Click Public projects in the top navigation bar.
- Locate the project and click Copy project in the lower right corner.
- In the pop-up window, you can name your copy of the project, select a billing group and decide whether this project will contain controlled data.
- Once you've customized the details, click Copy to copy the entire project.
You'll be redirected to the dashboard of your cloned project when it is ready, as shown below.
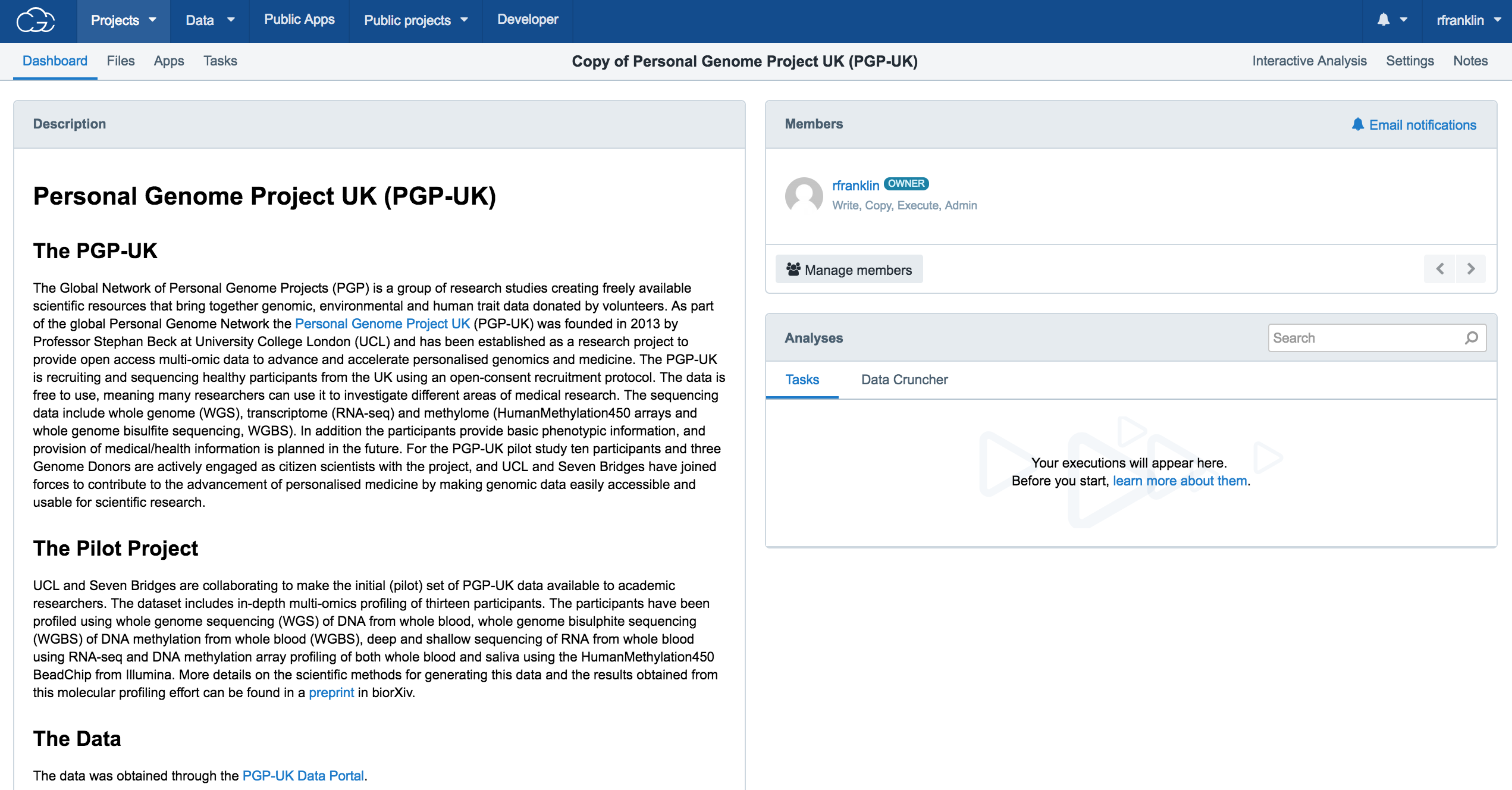
You can now use the data and workflows contained within this project to conduct your own analyses.
USE A SUBSET OF THE DATA
Instead of cloning the entire project, you can choose to select and copy a subset of the data.
- Access the PGP-UK public project (see above). You'll be taken to the project dashboard of the PGP-UK public project, as shown below.
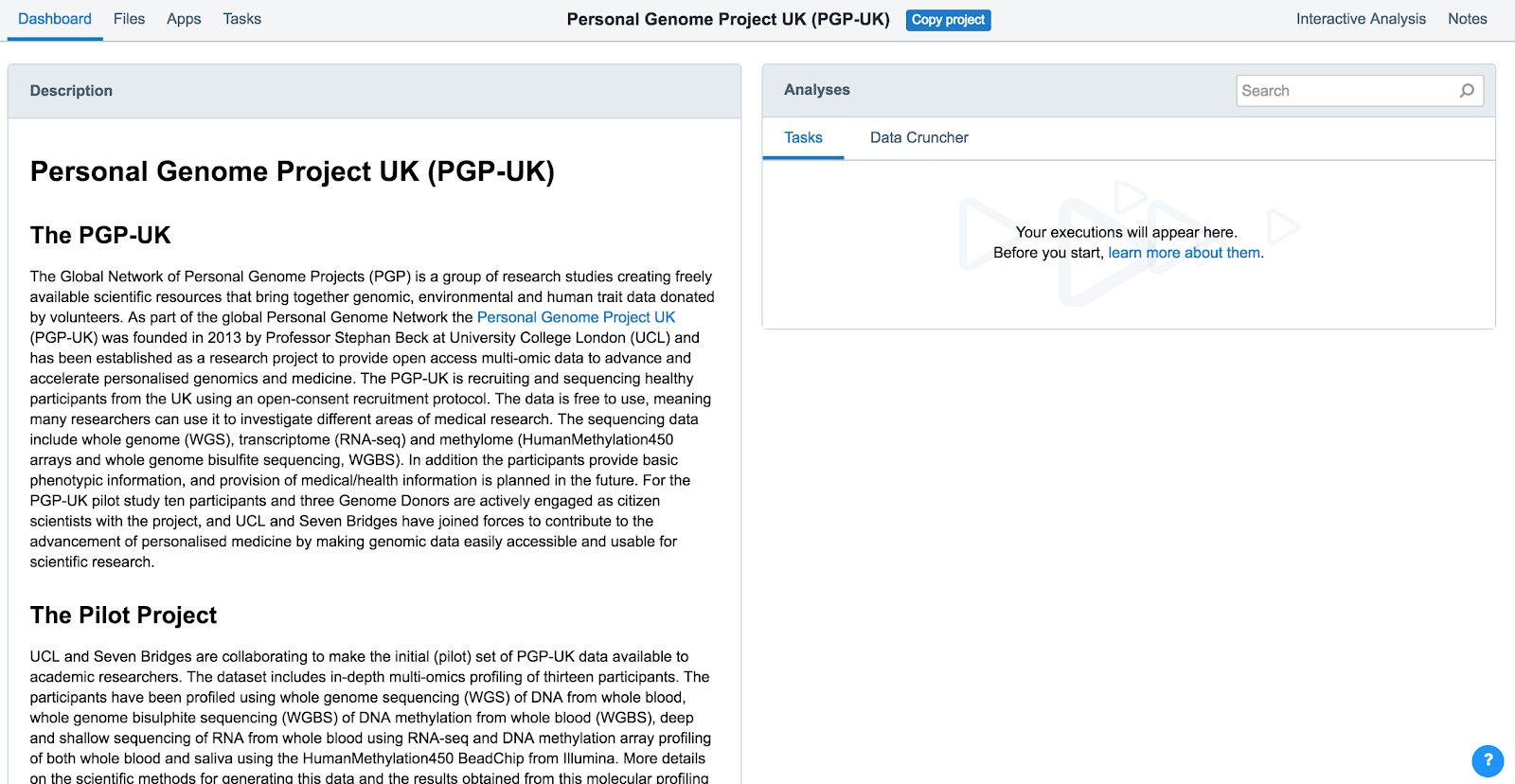
- Click Files in the upper lefthand corner. This will take you to the Files tab for the PGP-UK project, as shown below.
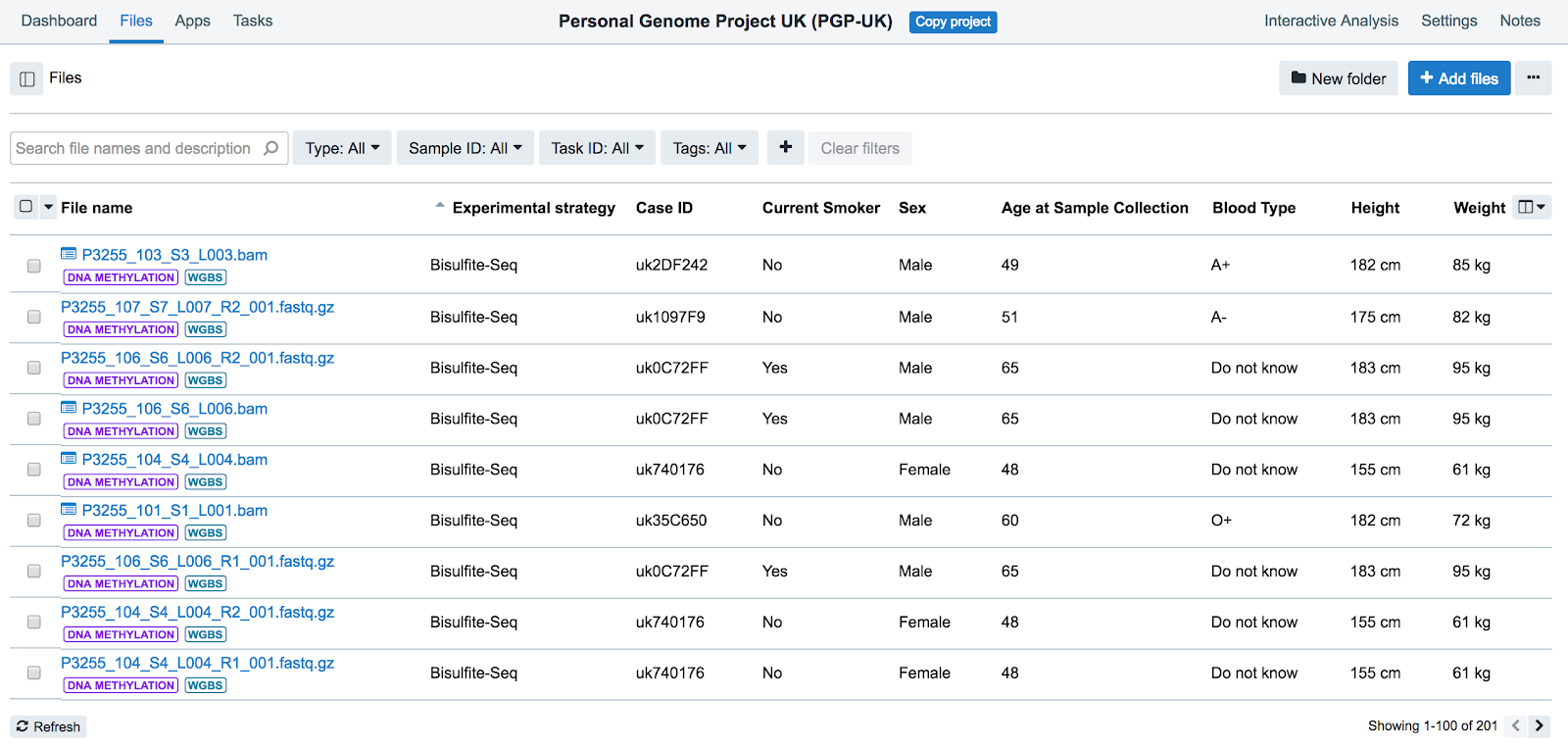
-
Filter or search for the desired files. You can filter by:
- Keywords - You can use the search bar at the top of the page to find files by entering the file name or notes associated with a file.
- Metadata fields - Next to the search bar, you will see drop-down menus for the available metadata fields. Use metadata fields such as Experimental strategy, File type, and Case ID to find the samples you’re interested in. Selecting a particular metadata value from one of these menus displays only files that match the value. For example, filter by RNA-Seq in the Experimental strategy field to only see RNA-Seq data. You can add additional drop-down menus to filter by other metadata fields by clicking the + icon.
-
You can choose specific files by selecting the corresponding checkbox in front of the file name.
-
Select as many files as you desire and click Copy to.
-
Select your desired project from the drop-down menu.
Now, you can start using the PGP-UK files you've added to your personal project in your own analysis.
Updated about 3 years ago
