CCLE data
ABOUT DATASETS > CCLE data
Seven Bridges is committed to providing CGC users with the most up-to-date version of the CCLE dataset that is available from the NCI Genomic Data Commons (GDC). In keeping with this commitment, the CGC transitioned from hosting the CGHub version of this dataset to the GDC Legacy Archive Data Release 11.0 version on July 10, 2018. As of this date, all files accessible via the Data Browser and the API correspond to files in the GDC Legacy Archive. Files that were added to individual projects before this date and are no longer represented in the new dataset version will no longer be accessible via those projects but may be obtainable from the GDC archive by contacting the GDC Help Desk. We look forward to continuing to collaborate with the GDC in the months ahead to ensure the timely availability through the CGC of new data releases for this dataset.
The Cancer Cell Line Encyclopedia (CCLE) contains Open Access sequencing data for nearly 1000 cancer cell line samples. The CCLE is made possible through a collaboration between the Broad Institute, the Novartis Institutes for Biomedical Research, and the Genomics Institute of the Novartis Research Foundation.
Data from the CCLE is available on the CGC via a public project, which acts as a repository for data as well as for examples of specific analyses and the tools you need to replicate these analyses. You can copy any CCLE data into your own projects, where you can analyze it. You don't need special access or authorization status to use the data available in the CCLE project.
The CCLE public project contains the following distribution of samples and files by experimental strategy.
The CCLE dataset is termed "legacy" in accordance with the distinction GDC makes between harmonized and non-harmonized datasets. Learn more about legacy datasets.
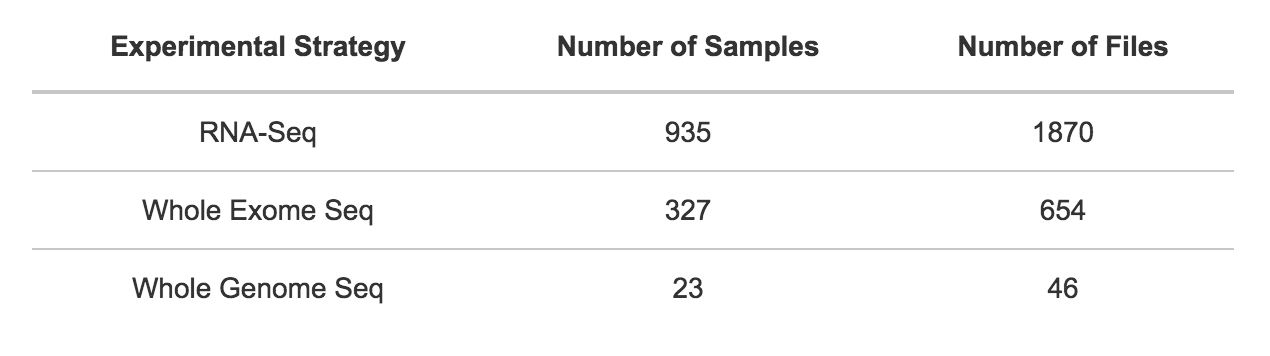
Browse and query the CCLE dataset. Or, access a repository of CCLE files via the CCLE public project.
Updated less than a minute ago
