Simons Genome Diversity Project (SGDP) dataset
Overview
The Simons Genome Diversity Project (SGDP) public project contains large Open Access files from the SGDP dataset which you can use on the CGC.
The SGDP dataset is made possible by the Simons Foundation. The dataset contains complete genome sequences from more than one hundred diverse human populations. It is the largest dataset of diverse, high quality human genome sequences ever reported. To represent as much anthropological, linguistic, and cultural diversity as possible, the dataset includes many deeply divergent human populations that are not well-represented in other datasets.
The SGDP public project contains Open Access whole genome sequencing data for 279 samples.
You don't need special access or authorization status to use the data in this project. In fact, any data you copy from this public project into your own projects will not count towards your storage.
The Simons Foundation asks that you please observe the Fort Lauderdale principles in your usage of SGDP data.
What's contained in the project?
The SGDP public project contains the following distribution of samples and files.

By geographical regions, the SGDP dataset is comprised of 44 Africans, 22 Native Americans, 27 Central Asians or Siberians, 47 East Asians, 25 Oceanians, 39 South Asians and 75 West Eurasians. Learn more about the metadata for the dataset.
Access the SGDP public project
- Click on Public projects from the top navigation bar.
- Select Simons Genome Diversity Project (SGDP) by clicking its title in the public projects gallery.
You'll be taken to the main dashboard of the SGDP public project.
Use the SGDP public project
All CGC users automatically have copy permissions for this project. This means that while you cannot upload data or tools to the project, you can copy the available data to your own projects on the CGC to execute analyses.
You have the options to:
- Copy the entire project - Start from the copied project and add apps to execute analyses on SGDP data.
- Select and copy a subset of the data to your own project - Use the selected data within your own analyses in your project.
Copy the entire project
- Click Public projects in the top navigation bar.
- Locate the project and click Copy project in the lower right corner.
- In the pop-up window, you can name your copy of the project, select a billing group and decide whether this project will contain controlled data.
- Once you've customized the details, click Copy to copy the entire project.
You'll be redirected to the dashboard of your cloned project when it is ready, as shown below. Add apps to conduct analyses on the data in your project.
Use a subset of the data
Instead of cloning the entire project, you can choose to select and copy a subset of the data.
- Access the SGDP public project by clicking Public projects in the top navigation bar and then selecting Simons Genome Diversity Project. You'll be taken to the project dashboard of the SGDP public project, as shown below.
- Click the Files tab in the upper righthand corner. This will take you to the Files page for the SGDP project, as shown below.
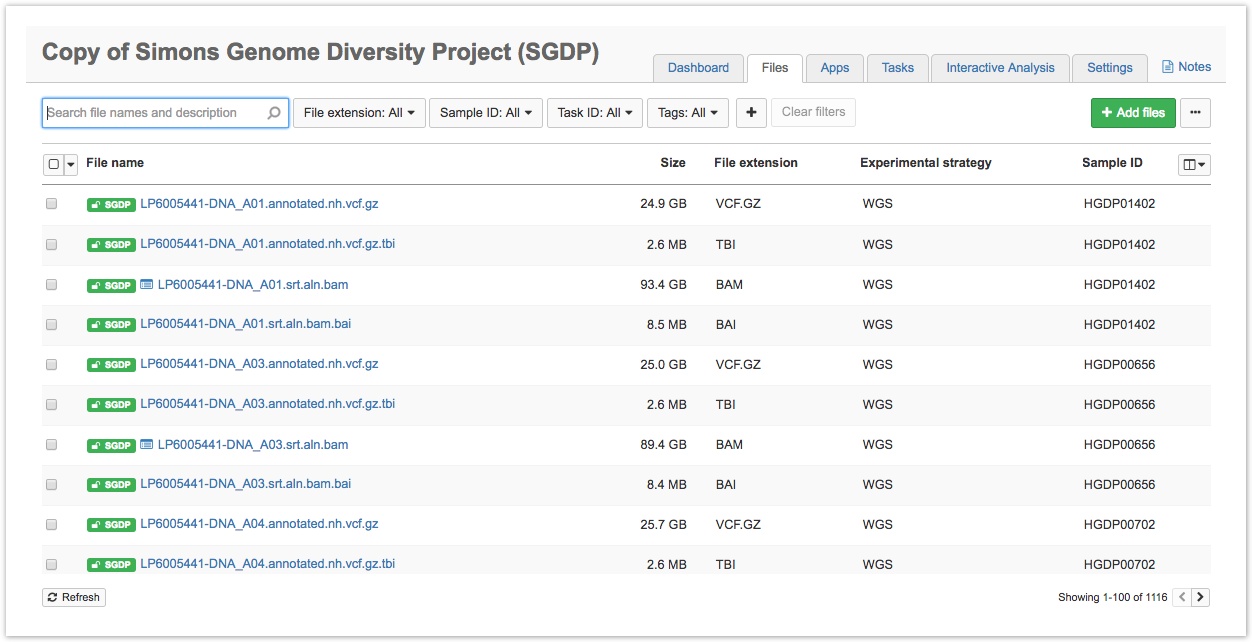
- Filter or search for the desired files. You can filter by:
- Keywords - You can use the search bar at the top of the page to find files by entering the file name or notes associated with a file.
- Metadata fields - Next to the search bar, you will see drop-down menus for the metadata fields Investigation, File extension, and Sample ID. Selecting a particular metadata value from one of these menus displays only files that match the value. For example, filter by SGDP-Australian in the Investigation field to only see samples from the Australian population. You can add additional drop-down menus to filter by other metadata fields by clicking the + icon.
- You can choose specific files by selecting the corresponding checkbox in front of the file name.
- Select as many files as you desire and click Copy to.
- Select your desired project from the drop-down menu.
Now, you can start using the SGDP files you've added to your personal project in your own analysis.
Updated about 3 years ago
